
Contact
How is it that cells manage to regulate their shape and organization during embryonic development in order to form the various tissues and diverse body plans seen in adult organisms?
Figure 1. Wildtype E8.5 mouse embryo was stained to detect Shroom3 (green) and F-actin (red). The neural plate was then imaged via confocal microscopy. 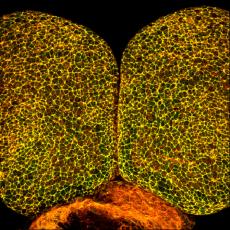
In most circumstances, cells utilize the coordinated efforts of signaling pathways, effector proteins, cytoskeletal networks, and contractile myosins to elicit the changes in cell morphology needed toform tissues and structures with beautiful and elaborate architecture. Understanding the regulation and integration of these cellular components, pathways, and networks in these fascinating processes is the main objective of the work in the Hildebrand lab. We use a variety of genetic, cellular, biochemical, molecular, and structural approaches to understand how cell and tissue morphology is regulated. We are currently studying the function and regulation of the Shroom family of proteins as a model to understand these processes.
Using numerous in vivo and in vitro model systems and approaches, we have shown that Shroom proteins are a family of actin-associated scaffolding molecules that control cellular architecture and tissue morphology during processes such as neural tube closure (Figure 1), kidney formation, vascularization (Figure 2), and Drosophila tissue morphogenesis (Figure 3).To date we have characterized the functions of vertebrate Shroom2, 3, and 4 and the ortholog of Shroom,from Drosophila melanogaster. It appears that all Shroom proteins control cell and tissue architecture by regulating the distribution of contractile actomyosin networks.
Figure 2. Shroom2 (green) and PECAM (red) expression in the vasculature network of the mouse yolk sac. 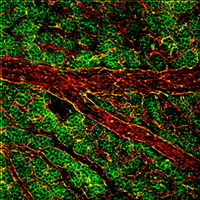
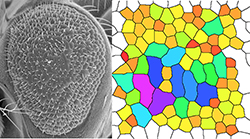

Figure 3. Rough eye phenotype (left) and quantification (right) caused by Shroom expression in the Drosophila eye imaginal disc.
- Nisha Holay, undergraduate researcher
- Jad Hilal, undergraduate researcher
- Serena Tally, undergraduate researcher
Zapata J, Moretto E, Hannan S, Murru L, Longatt
Odenwald MA, Choi W, Buckley A, Shashikanth N,
Zalewski JK, Heber S, Mo JH, O'Conor K, Hildebr
Zalewski JK, Mo JH, Heber S, Heroux A, Gardner
Grego-Bessa J, Hildebrand JD, Anderson KV (2015
McGreevy EM, Vijayraghavan D, Davidson LA, Hild
Das D, Zalewski JK, Mohan S, Plageman TF, VanDe
Lang RA, Herman K, Reynolds AB, Hildebrand JD,
Mohan, S., Das, D., Bauer, RJ, Heroux, A., Zale
Mohan S, Rizaldy R, Das D, Bauer RJ, Heroux A,
Plageman TF Jr, Chauhan BK, Yang C, Jaudon F, Shang X, Zheng Y, Lou M, Debant A, Hildebrand JD, L
Grosse AS, Pressprich MF, Curley LB, Hamilton KL, Margolis B, Hildebrand JD, Gumucio DL. (2011) C
Farber, M.J., R. Rizaldy, and J.D. Hildebrand (2011) Shroom2 regulates contractility to control e
Mo, D., B.A. Potter, C.A. Bertrand, J.D. Hildebrand, J.R. Bruns, and O.A. Weisz (2010) Nucleofect
Bolinger, C., L. Zasadil, R. Rizaldy, and J.D. Hildebrand (2010) Specific isoforms of Drosoph
Plageman TF, J.r, M.I. Chung, M. Lou, A.N. Smith, J.D. Hildebrand, J.B. Wallingford, and R.A. Lan
Yoder, M., and J.D. Hildebrand (2007) Shroom4 (KIAA1202) is an actin-associated protein implicate
Hagens, O., A. Ballabio, V. Kalscheuer, J.P. Kraehenbuhl, M.V. Schiaffino, P. Smith, O. Staub, J.
Fairbank, P.D., C. Lee, A. Ellis, J.D. Hildebrand, J.M. Gross, and J.B. Wallingford (2006) Shroom
Dietz, M.L., T.M. Bernaciak, F. Vendetti, and J.D. Hildebrand (2006) Differential actin-dependent
Hildebrand, J.D. (2005) CtBP proteins in vertebrate development. Pp in CtBP Family Proteins
Hildebrand, J.D. (2005) Shroom regulates epithelial cell shape via the apical positioning of an a
Haigo, S.L., J.D. Hildebrand, R.M. Harland, and J.B. Wallingford (2003) Shroom induces apical con
Zhang, Q., Y. Yoshimatsu, J. Hildebrand, S.M. Frisch, and R.H. Goodman (2003) Homeodomain Interac
Lin, X., B. Sun, M. Liang, Y.Y. Liang, A. Gast, J. Hildebrand, F.C. Brunicardi, F. Melchior, and
Grooteclaes, M., Q. Deveraux, J. Hildebrand, Q. Zhang, R.H. Goodman, and S.M. Frisch (2003) C-ter
Hildebrand, J.D., and P. Soriano (2002) Overlapping and unique roles for C-terminal binding prote
