
Contact
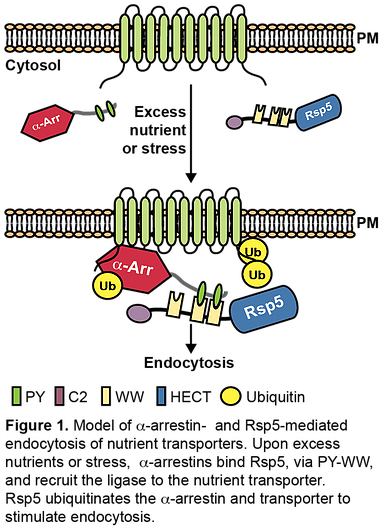 Nearly half of all prescription drugs alter G-protein coupled receptor (GPCR) signaling, including treatments for asthma, hypertension, neurodegenerative disorders and depression. β-arrestins are critical regulators of GPCRs: they act as trafficking adaptors to control GPCR endocytosis and impede G-protein signaling. β-arrestins are themselves therapeutic targets, highlighting the clinical importance of understanding arrestin function. However, β-arrestins are only a small branch of the larger arrestin family that includes the widely-conserved but functionally uncharacterized α-arrestins, the primary focus of our research. Our work has shown that α-arrestins, like β-arrestins, regulate GPCR signaling, but also operate in unexpected trafficking pathways, including endosomal recycling and clathrin-independent endocytosis. Using Saccharomyces cerevisiae as a model, our research has shown that α-arrestins interact with signaling regulators, including kinases and phosphatases, to selectively direct trafficking of specific transmembrane cargo proteins, presumably by promoting their ubiquitination and packaging into vesicles (Figure 1), and have begun to define the molecular mechanisms underlying α-arrestin-mediated trafficking. All of the α-arrestin-interacting partners identified in yeast are conserved. Our research will apply insights gained in yeast to initiate studies on the relatively unstudied mammalian α-arrestins.
Nearly half of all prescription drugs alter G-protein coupled receptor (GPCR) signaling, including treatments for asthma, hypertension, neurodegenerative disorders and depression. β-arrestins are critical regulators of GPCRs: they act as trafficking adaptors to control GPCR endocytosis and impede G-protein signaling. β-arrestins are themselves therapeutic targets, highlighting the clinical importance of understanding arrestin function. However, β-arrestins are only a small branch of the larger arrestin family that includes the widely-conserved but functionally uncharacterized α-arrestins, the primary focus of our research. Our work has shown that α-arrestins, like β-arrestins, regulate GPCR signaling, but also operate in unexpected trafficking pathways, including endosomal recycling and clathrin-independent endocytosis. Using Saccharomyces cerevisiae as a model, our research has shown that α-arrestins interact with signaling regulators, including kinases and phosphatases, to selectively direct trafficking of specific transmembrane cargo proteins, presumably by promoting their ubiquitination and packaging into vesicles (Figure 1), and have begun to define the molecular mechanisms underlying α-arrestin-mediated trafficking. All of the α-arrestin-interacting partners identified in yeast are conserved. Our research will apply insights gained in yeast to initiate studies on the relatively unstudied mammalian α-arrestins.
The study of α-arrestins is in its infancy. There are still many unanswered questions about arrestin biology: What are the initial signaling cues that regulate α-arrestin trafficking? How are specific cargo proteins recognized? How does the arrestin-cargo interaction direct a protein cargo to its final destination? Our research employs molecular, biochemical, genetic and advanced microscopy methods to address these fundamental questions about arrestin function in yeast to expand our understanding of GPCR signaling and protein trafficking.
- Allyson O'Donnell, Assistant Professor
- Annette Chiang, Research Associate
- Mitchell Lesko, Graduate Student
- Elif Filiztekin, Graduate Student
- Carly Houghton, Graduate Student
- Nikolai Makhov, Rotating Graduate Student
- Prathiksha Sivakumar, Undergraduate Researcher
- Ruiqi He, Undergraduate Researcher
- Charlotte Qi, Undergraduate Researcher
- Amalia Kamon, Undergraduate Researcher
- Emma Bocquillon, Undergraduate Researcher
O’Donnell, A.F. (2022) A second chance at yeast
Bowman, R.W., S. Davis, E.M. Jordahl, K.Chera,
Hager, N.A., C.K. McAtee, M.A. Lesko and A.F. O’Do
Hellmann, E.*, J. Walker*, M.A. Lesko*, D. Chandra
Bowman, R.W.*, E.M. Jordahl*, S. Davis*, S. Hed
Robinson, B., S. Hawbaker, A. Chiang, E. Jordahl,
Schmidt, M.C.¶ and A.F. O’Donnell¶ (2020) ‘Sugarco
Vakirlis, N., B. Hsu, O. Acar, N. Castilho Coelho,
Soncini, S.R., D.G. Chandrashekarappa, D.A. August
O’Donnell, A.F.‡ and M.S. Schmidt‡ (2019) AMPK-
Hager, N.A., C.J. Krasowski, T.D. Mackie, A.R.
O’Donnell, A.F. and Schmidt, M.C. (2019) Helping d
Mackie, T.D., B.Y. Kim, A.R. Subramanya, D.J. B
Chandrashekarappa, D.G., R. R. McCartney, A.F.
Prosser, D.C., K. Wrasman, T. K. Woodard, A. F.
Prosser, D.C., A.E. Pannunzio, J. Brodsky, J. T
O’Donnell, A.F., R. R. McCartney, D. G.
C.G. Alvaro, A.F. O’Donnell*, D.C. Pross
Hecht, K.A., A.F. O’Donnell, and J.L. Brodsky.
O’Donnell, A.F., L. Huang, J. Thorner, a
O’Donnell, A.F. (2012) The running of th
Stevens, J.R., A.F. O’Donnell, T.E. Perr
Minear, S., A.F. O’Donnell, G. Giaever,
Piña, F.J., A.F. O’Donnell, S. Pagant, H
O’Donnell, A.F., A. Apffel, R.G. Gardner
O’Donnell, A.F., J.R. Stevens, R. Kepkay
O’Donnell, A.F., N.K. Brewster, J. Kurni
O’Donnell, A.F., S. Tiong, D. Nash, and
Dr. Allyson O'Donnell joined the department in 2018. Allyson is also a member of the Center for Protein Conformational Diseases at the University of Pittsburgh ( http://www.proteindiseasecenter.pitt.edu/about-our-center), The Pittsburgh Center for Evolutionary Biology and Medicine (https://www.cebam.pitt.edu/), and the American Society for Cell Biology. Allyson serves as a mentor for the Science Outreach Program at Taylor Allderdice High School.
Dr. O'Donnell received her B.S. degree in Biochemistry and M.S. degree in Biology from the University of New Brunswick (Canada). During her Masters’ thesis work she identified genes needed for de novo purine biosynthesis in Drosophila melanogaster and characterized the developmental defects associated with mutations in these genes. Dr. O'Donnell went on to receive her Ph.D. in Biochemistry & Molecular Biology from Dalhousie University (Canada) where she studied the role of the FACT histone chaperone complex in chromatin remodeling.
During her post-doctoral research at Stanford University and the University of California, Berkeley Dr. O'Donnell began her research on a previously unstudied class of protein trafficking adaptors, now referred to as the α-arrestins. Her research has shown that α-arrestins regulate trafficking of G-protein coupled receptors, but also operate in unexpected trafficking pathways, including endosomal recycling and clathrin-independent endocytosis. Using Saccharomyces cerevisiae as a model, she has identified α-arrestin interactions with signaling regulators, cargos and vesicle coat proteins, and has begun to define the molecular mechanisms underlying α-arrestin-mediated trafficking. Her research applies insights gained in yeast to target studies on the relatively unstudied mammalian α-arrestins.
