
Contact
The development of multicellular animals requires precise regulation of gene activity and cell behavior to control patterns of cell differentiation and final size and shape.
We use the fruit fly, Drosophila melanogaster, to answer questions such as: why is a gene switched off in one cell but not in another, and why are appendages generally long and thin rather than short and fat? Drosophila offers a wide range of genetic and molecular approaches to study development and because genetic pathways are shared with other animals the results from these studies help us to understand similar phenomena in other species, even to revealing mechanisms involved in human development and disease, such as cancer.
Repression
Not all of the genes in a cell are being actively transcribed; cells have to switch genes on and off to control their differentiation and to respond to various stimuli. Genes can be silenced by repressors - regulatory transcription factors - which bind to specific sequences adjacent coding regions and use various mechanisms to negatively influence transcription. Most repressors, such as the fly protein, Brinker, recruit secondary proteins, corepressors, including Groucho and CtBP. We are investigating why repressors such as Brinker need to recruit multiple corepressors and how these corepressors function to provide repressive activity. Recent studies indicate that Brk needs to recruit more than one corepressor because its primary one, Groucho, is unavailable in some tissues where CtBP acts as a backup.
Gradients and thresholds
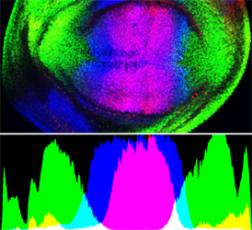
In developing tissues, precise patterns of gene expression are established that presage the final pattern of differentiation. A common mechanism used to achieve this is the generation of a transcription factor gradient and the activation and repression of different target genes above distinct thresholds. For example, in the developing fly wing Brinker is expressed in symmetrical lateral-to-medial gradients (Fig. 1) and target genes sal and omb are expressed in overlapping, nested patterns with the omb domain wider than sal because omb is less sensitive to repression by Brk. We are studying how the Brk gradient is established and what determines how sensitive a gene is to repression by Brk.
Size and shape
Each organ or appendage, such as the fly wing, has to achieve the correct size and shape to function properly. The cellular mechanisms operating to generate the elongated morphology of the wing can be revealed by mutations which disrupt its shape and are beginning to show that regulating patterns of growth, cell movement and cell shape are key to morphogenesis of this appendage. We are characterizing several mutants, such as narrow (Fig. 2), to determine the relative importance of these processes at different stages of development.

